Yesterday morning, my last day at the Lake Arrowhead Microbial Genomics Conference, I saw a tweet from Holly Bik (@Dr_Bik) about a talk she was attending at a Phenomics conference about the sociology of Amazon’s Mechanical Turk Web Service. What is Mechanical Turk, you may ask? Well, what’s really funny is that just minutes before I had answered that question for Ben Tully (@phantomBugs) in describing how I used it to help with my dissertation research.
Mechanical Turk allows you to crowd source little tasks that are easy for humans, but not computers. For example, if you need to write a short caption for 1000 photos at about 1 minute per photo, that would take you about 2 full days of work. Or, you can upload those photos to Mechanical Turk, along with some instructions about how to write each caption. Each photo becomes a little job, sent out to all the workers on Mechanical Turk. You offer to pay $.03 per job, and then you sit back with a glass of wine and watch the World Series of Poker. Some hours later, all of your work is done, and you did none of it. Sure, you are out $30, but hopefully your time is worth more than $15/day.
You are provided with the worker ID for each job. You can spot-check each worker’s work and if you do not like it, you can reject all of their work, do not pay them, and then those photos go back into the work queue
So, how did I use this for my dissertation research? I wanted to look at environmental correlates of horizontal gene transfer (HGT). HGT is the exchange of DNA between different species, and it is fairly common among microbes. One potentially important mechanism of DNA transfer is the uptake by a cell of DNA that is floating about freely in the environment (transformation). If the environment is inhospitable to the DNA molecule, then the probability of transfer by transformation should be quite low. For example, and in particular, I wanted to ask whether organisms that live in very low pH environments experience a lower incidence of HGT.
I can predict the incidence of HGT for an organism directly from the genome sequence, so all I need is to find out at what pH that organism grows. That should be straightforward because every time a genome is submitted to a public database, the submitter will include all of the associated environmental data (or metadata) that is available, and since that submitter grew the organism in culture in the laboratory, he or she must know at which pH it best grows. Right?
Now, because I am who I am, I want to do this analysis in a phylogenetic context, using some Phylogenetic Comparative Method (I should talk about this more in another post.) In particular, I opted to use Felsenstein’s Independent Contrast method as implemented in Phylocom. I built a reference phylogeny for ~800 bacteria and archaea which have genome sequences available (this part is currently in revision) and then I “looked up” the optimum growth pH for each of them. This look up process should be straightforward, too, because the data are submitted to a searchable database, like at NCBI or the JGI’s IMG. Right?
Well, nothing that should be straightforward ever is in my academic life. I was able to get the pH data from the IMG by grabbing all of the webages with the metadata on them and parsing them with a little perl script. But when I did that, I only got pH data for ~100 of the ~800 organisms in my reference phylogeny. For any given organism, if I spent a couple of minutes poking around in the literature, I could easily find the optimum growth pH, so it’s not like it’s not out there. But, I couldn’t automate the “poking around” process, and after spending two full days of work, I had only retrieved pH data for about 200 additional organisms. Because I was down to the wire in terms of a dissertation submission deadline, and because I consider my time fairly valuable, I just couldn’t bring myself to keep at it.
I was complaining vociferously about those ~600 people who couldn’t be bothered to spend their couple of minutes to include pH data with their genome submissions. Russell Neches (@ryneches), who seemed to really get a kick out of my uncharacteristic vociferousness, suggested that Mechanical Turk might work for me.
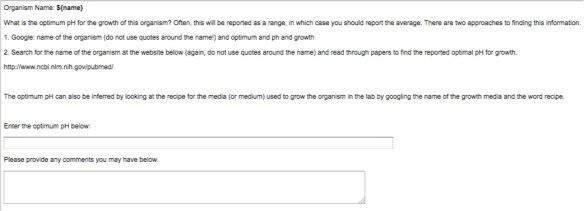
I created a job (called a HIT): “Given the name of an organism, report it’s optimum growth pH”
My job looked like this:
In retrospect, maybe I could have come up with some better instructions, but I think for most things, these should work. I paid $.08 per job, so I spent $40. I rejected a lot of work because I got answers like: 37.5 or comments like “I couldn’t find it.” There was one worker who commented, “This was a cool HIT.” And that worker did a lot of jobs and seemed to do good work. I focused my spot-checking on values at the extremes, since most things live at a neutral pH.
But, here’s what I found: If you follow the first approach that I suggest and google the organism name+optimum+pH+growth, sometimes, you would see something that looked right in the google search results, but was actually referring to enzyme activity rather than growth conditions.
I can tell that this is not the information I’m looking for, but I don’t really expect a Mechanical Turk worker to be so discerning, especially not for $.08. There were a few other common errors that I think it would have been difficult to avoid, even with more clear instructions. I could have submitted some of the jobs in duplicate so that I could check the workers against each other, but I suspect that this type of mistake would be made by anyone without a fair degree of specialized knowledge on the subject (who is not likely to be doing this sort of work.)
So, in the end, I ended up spending a lot of time double-checking the results. I don’t know if I saved any time by doing it this way, but it was FUN! I will definitely keep it in the back of my mind, and hope to be able to use it again someday!